AWS Machine Learning Blog
Exploring summarization options for Healthcare with Amazon SageMaker
In today’s rapidly evolving healthcare landscape, doctors are faced with vast amounts of clinical data from various sources, such as caregiver notes, electronic health records, and imaging reports. This wealth of information, while essential for patient care, can also be overwhelming and time-consuming for medical professionals to sift through and analyze. Efficiently summarizing and extracting insights from this data is crucial for better patient care and decision-making. Summarized patient information can be useful to a number of downstream processes like data aggregation, effectively coding patients, or grouping patients with similar diagnoses for review.
Artificial intelligence (AI) and machine learning (ML) models have shown great promise in addressing these challenges. Models can be trained to analyze and interpret large volumes of text data, effectively condensing information into concise summaries. By automating the summarization process, doctors can quickly gain access to relevant information, allowing them to focus on patient care and make more informed decisions. See the following case study to learn more about a real-world use case.
Amazon SageMaker, a fully managed ML service, provides an ideal platform for hosting and implementing various AI/ML-based summarization models and approaches. In this post, we explore different options for implementing summarization techniques on SageMaker, including using Amazon SageMaker JumpStart foundation models, fine-tuning pre-trained models from Hugging Face, and building custom summarization models. We also discuss the pros and cons of each approach, enabling healthcare professionals to choose the most suitable solution for generating concise and accurate summaries of complex clinical data.
Two important terms to know before we begin: pre-trained and fine-tuning. A pre-trained or foundation model is one that has been built and trained on a large corpus of data, typically for general language knowledge. Fine-tuning is the process by which a pre-trained model is given another more domain-specific dataset in order to enhance its performance on a specific task. In a healthcare setting, this would mean giving the model some data including phrases and terminology pertaining specifically to patient care.
Build custom summarization models on SageMaker
Though the most high-effort approach, some organizations might prefer to build custom summarization models on SageMaker from scratch. This approach requires more in-depth knowledge of AI/ML models and may involve creating a model architecture from scratch or adapting existing models to suit specific needs. Building custom models can offer greater flexibility and control over the summarization process, but also requires more time and resources compared to approaches that start from pre-trained models. It’s essential to weigh the benefits and drawbacks of this option carefully before proceeding, because it may not be suitable for all use cases.
SageMaker JumpStart foundation models
A great option for implementing summarization on SageMaker is using JumpStart foundation models. These models, developed by leading AI research organizations, offer a range of pre-trained language models optimized for various tasks, including text summarization. SageMaker JumpStart provides two types of foundation models: proprietary models and open-source models. SageMaker JumpStart also provides HIPAA eligibility, making it useful for healthcare workloads. It is ultimately up to the customer to ensure compliance, so be sure to take the appropriate steps. See Architecting for HIPAA Security and Compliance on Amazon Web Services for more details.
Proprietary foundation models
Proprietary models, such as Jurassic models from AI21 and the Cohere Generate model from Cohere, can be discovered through SageMaker JumpStart on the AWS Management Console and are currently under preview. Utilizing proprietary models for summarization is ideal when you don’t need to fine-tune your model on custom data. This offers an easy-to-use, out-of-the-box solution that can meet your summarization requirements with minimal configuration. By using the capabilities of these pre-trained models, you can save time and resources that would otherwise be spent on training and fine-tuning a custom model. Furthermore, proprietary models typically come with user-friendly APIs and SDKs, streamlining the integration process with your existing systems and applications. If your summarization needs can be met by pre-trained proprietary models without requiring specific customization or fine-tuning, they offer a convenient, cost-effective, and efficient solution for your text summarization tasks. Because these models are not trained specifically for healthcare use cases, quality can’t be guaranteed for medical language out of the box without fine-tuning.
Jurassic-2 Grande Instruct is a large language model (LLM) by AI21 Labs, optimized for natural language instructions and applicable to various language tasks. It offers an easy-to-use API and Python SDK, balancing quality and affordability. Popular uses include generating marketing copy, powering chatbots, and text summarization.
On the SageMaker console, navigate to SageMaker JumpStart, find the AI21 Jurassic-2 Grande Instruct model, and choose Try out model.

If you want to deploy the model to a SageMaker endpoint that you manage, you can follow the steps in this sample notebook, which shows you how to deploy Jurassic-2 Large using SageMaker.
Open-source foundation models
Open-source models include FLAN T5, Bloom, and GPT-2 models that can be discovered through SageMaker JumpStart in the Amazon SageMaker Studio UI, SageMaker JumpStart on the SageMaker console, and SageMaker JumpStart APIs. These models can be fine-tuned and deployed to endpoints under your AWS account, giving you full ownership of model weights and script codes.
Flan-T5 XL is a powerful and versatile model designed for a wide range of language tasks. By fine-tuning the model with your domain-specific data, you can optimize its performance for your particular use case, such as text summarization or any other NLP task. For details on how to fine-tune Flan-T5 XL using the SageMaker Studio UI, refer to Instruction fine-tuning for FLAN T5 XL with Amazon SageMaker Jumpstart.
Fine-tuning pre-trained models with Hugging Face on SageMaker
One of the most popular options for implementing summarization on SageMaker is fine-tuning pre-trained models using the Hugging Face Transformers library. Hugging Face provides a wide range of pre-trained transformer models specifically designed for various natural language processing (NLP) tasks, including text summarization. With the Hugging Face Transformers library, you can easily fine-tune these pre-trained models on your domain-specific data using SageMaker. This approach has several advantages, such as faster training times, better performance on specific domains, and easier model packaging and deployment using built-in SageMaker tools and services. If you’re unable to find a suitable model in SageMaker JumpStart, you can choose any model offered by Hugging Face and fine-tune it using SageMaker.
To start working with a model to learn about the capabilities of ML, all you need to do is open SageMaker Studio, find a pre-trained model you want to use in the Hugging Face Model Hub, and choose SageMaker as your deployment method. Hugging Face will give you the code to copy, paste, and run in your notebook. It’s as easy as that! No ML engineering experience required.

The Hugging Face Transformers library enables builders to operate on the pre-trained models and do advanced tasks like fine-tuning, which we explore in the following sections.
Provision resources
Before we can begin, we need to provision a notebook. For instructions, refer to Steps 1 and 2 in Build and Train a Machine Learning Model Locally. For this example, we used the settings shown in the following screenshot.
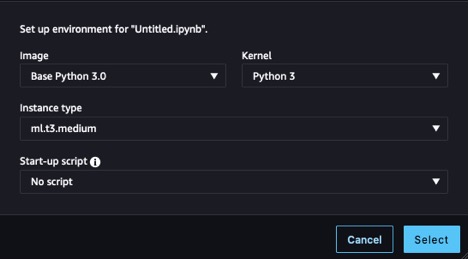
We also need to create an Amazon Simple Storage Service (Amazon S3) bucket to store the training data and training artifacts. For instructions, refer to Creating a bucket.
Prepare the dataset
To fine-tune our model to have better domain knowledge, we need to get data suitable for the task. When training for an enterprise use case, you’ll need to go through a number of data engineering tasks to prepare your own data to be ready for training. Those tasks are outside the scope of this post. For this example, we’ve generated some synthetic data to emulate nursing notes and stored it in Amazon S3. Storing our data in Amazon S3 enables us to architect our workloads for HIPAA compliance. We start by getting those notes and loading them on the instance where our notebook is running:
The notes are composed of a column containing the full entry, note, and a column containing a shortened version exemplifying what our desired output should be, summary. The purpose of using this dataset is to improve our model’s biological and medical vocabulary so that it’s more attuned to summarizing in a healthcare context, called domain fine-tuning, and show our model how to structure its summarized output. In some summarization cases, we may want to create an abstract out of an article or a one-line synopsis of a review, but in this case, we’re trying to get our model to output an abbreviated version of the symptoms and actions taken for a patient so far.
Load the model
The model we use as our foundation is a version of Google’s Pegasus, made available in the Hugging Face Hub, called pegasus-xsum. It’s already pre-trained for summarization, so our fine-tuning process can focus on extending its domain knowledge. Modifying the task our model runs is a different type of fine-tuning not covered in this post. The Transformer library supplies us with a class to load the model definition from our model_checkpoint: google/pegasus-xsum. This will load the model from the hub and instantiate it in our notebook so we can use it later on. Because pegasus-xsum is a sequence-to-sequence model, we want to use the Seq2Seq type of the AutoModel class:
Now that we have our model, it’s time to put our attention to the other components that will enable us to run our training loop.
Create a tokenizer
The first of these components is the tokenizer. Tokenization is the process by which words from the input data are transformed into numerical representations that our model can understand. Again, the Transformer library provides a class for us to load a tokenizer definition from the same checkpoint we used to instantiate the model:
With this tokenizer object, we can create a preprocessing function and map it onto our dataset to give us tokens ready to be fed into the model. Finally, we format the tokenized output and remove the columns containing our original text, because the model will not be able to interpret them. Now we’re left with a tokenized input ready to be fed into the model. See the following code:
With our data tokenized and our model instantiated, we’re almost ready to run a training loop. The next components we want to create are the data collator and the optimizer. The data collator is another class provided by Hugging Face through the Transformers library, which we use to create batches of our tokenized data for training. We can easily build this using the tokenizer and model objects we already have just by finding the corresponding class type we’ve used previously for our model (Seq2Seq) for the collator class. The optimizer’s function is to maintain the training state and update the parameters based on our training loss as we work through the loop. To create an optimizer, we can import the optim package from the torch module, where a number of optimization algorithms are available. Some common ones you may have encountered before are Stochastic Gradient Descent and Adam, the latter of the which is applied in our example. Adam’s constructor takes in the model parameters and the parameterized learning rate for the given training run. See the following code:
The last steps before we can begin training are to build the accelerator and the learning rate scheduler. The accelerator comes from a different library (we’ve been primarily using Transformers) produced by Hugging Face, aptly named Accelerate, and will abstract away logic required to manage devices during training (using multiple GPUs for example). For the final component, we revisit the ever-useful Transformers library to implement our learning rate scheduler. By specifying the scheduler type, the total number of training steps in our loop, and the previously created optimizer, the get_scheduler function returns an object that enables us to adjust our initial learning rate throughout the training process:
We’re now fully set up for training! Let’s set up a training job, starting by instantiating the training_args using the Transformers library and choosing parameter values. We can pass these, along with our other prepared components and dataset, directly to the trainer and start training, as shown in the following code. Depending on the size of your dataset and chosen parameters, this may take a significant amount of time.
Package the model for inference
After training has been run, the model object is ready to be used for inference. As a best practice, let’s save our work for future use. We need to create our model artifacts, zip them together, and upload our tarball to Amazon S3 for storage. To prepare our model for zipping, we need to unwrap the now fine-tuned model, then save the model binary and associated config files. We also need to save our tokenizer to the same directory that we saved our model artifacts to so it is available when we use the model for inference. Our model_dir folder should now look something like the following code:
All that’s left is to run a tar command to zip up our directory and upload the tar.gz file to Amazon S3:
Our newly fine-tuned model is now ready and available to be used for inference.
Perform inference
To use this model artifact for inference, open a new file and use the following code, modifying the model_data parameter to fit your artifact save location in Amazon S3. The HuggingFaceModel constructor will rebuild our model from the checkpoint we saved to model.tar.gz, which we can then deploy for inference using the deploy method. Deploying the endpoint will take a few minutes.
After the endpoint is deployed, we can use the predictor we’ve created to test it. Pass the predict method a data payload and run the cell, and you’ll get the response from your fine-tuned model:
To see the benefit of fine-tuning a model, let’s do a quick test. The following table includes a prompt and the results of passing that prompt to the model before and after fine-tuning.
| Prompt | Response with No Fine-Tuning | Response with Fine-Tuning |
| Summarize the symptoms that the patient is experiencing. Patient is a 45 year old male with complaints of substernal chest pain radiating to the left arm. Pain is sudden onset while he was doing yard work, associated with mild shortness of breath and diaphoresis. On arrival patient’s heart rate was 120, respiratory rate 24, blood pressure 170/95. 12 lead electrocardiogram done on arrival to the emergency department and three sublingual nitroglycerin administered without relief of chest pain. Electrocardiogram shows ST elevation in anterior leads demonstrating acute anterior myocardial infarction. We have contacted cardiac catheterization lab and prepping for cardiac catheterization by cardiologist. | We present a case of acute myocardial infarction. | Chest pain, anterior MI, PCI. |
As you can see, our fine-tuned model uses health terminology differently, and we’ve been able to change the structure of the response to fit our purposes. Note that results are dependent on your dataset and the design choices made during training. Your version of the model could offer very different results.
Clean up
When you’re finished with your SageMaker notebook, be sure to shut it down to avoid costs from long-running resources. Note that shutting down the instance will cause you to lose any data stored in the instance’s ephemeral memory, so you should save all your work to persistent storage before cleanup. You will also need to go to the Endpoints page on the SageMaker console and delete any endpoints deployed for inference. To remove all artifacts, you also need to go to the Amazon S3 console to delete files uploaded to your bucket.
Conclusion
In this post, we explored various options for implementing text summarization techniques on SageMaker to help healthcare professionals efficiently process and extract insights from vast amounts of clinical data. We discussed using SageMaker Jumpstart foundation models, fine-tuning pre-trained models from Hugging Face, and building custom summarization models. Each approach has its own advantages and drawbacks, catering to different needs and requirements.
Building custom summarization models on SageMaker allows for lots of flexibility and control but requires more time and resources than using pre-trained models. SageMaker Jumpstart foundation models provide an easy-to-use and cost-effective solution for organizations that don’t require specific customization or fine-tuning, as well as some options for simplified fine-tuning. Fine-tuning pre-trained models from Hugging Face offers faster training times, better domain-specific performance, and seamless integration with SageMaker tools and services across a broad catalog of models, but it requires some implementation effort. At the time of writing this post, Amazon has announced another option, Amazon Bedrock, which will offer summarization capabilities in an even more managed environment.
By understanding the pros and cons of each approach, healthcare professionals and organizations can make informed decisions on the most suitable solution for generating concise and accurate summaries of complex clinical data. Ultimately, using AI/ML-based summarization models on SageMaker can significantly enhance patient care and decision-making by enabling medical professionals to quickly access relevant information and focus on providing quality care.
Resources
For the full script discussed in this post and some sample data, refer to the GitHub repo. For more information on how to run ML workloads on AWS, see the following resources:
- Hugging Face on Amazon SageMaker Workshop
- Hugging Face Transformers Amazon SageMaker Examples
- Technology Innovation Institute trains the state-of-the-art Falcon LLM 40B foundation model on Amazon SageMaker
- Training large language models on Amazon SageMaker: Best practices
- How Forethought saves over 66% in costs for generative AI models using Amazon SageMaker
About the authors
 Cody Collins is a New York based Solutions Architect at Amazon Web Services. He works with ISV customers to build industry leading solutions in the cloud. He has successfully delivered complex projects for diverse industries, optimizing efficiency and scalability. In his spare time, he enjoys reading, traveling, and training jiu jitsu.
Cody Collins is a New York based Solutions Architect at Amazon Web Services. He works with ISV customers to build industry leading solutions in the cloud. He has successfully delivered complex projects for diverse industries, optimizing efficiency and scalability. In his spare time, he enjoys reading, traveling, and training jiu jitsu.
 Ameer Hakme is an AWS Solutions Architect residing in Pennsylvania. His professional focus involves collaborating with Independent software vendors throughout the Northeast, guiding them in designing and constructing scalable, state-of-the-art platforms on the AWS Cloud.
Ameer Hakme is an AWS Solutions Architect residing in Pennsylvania. His professional focus involves collaborating with Independent software vendors throughout the Northeast, guiding them in designing and constructing scalable, state-of-the-art platforms on the AWS Cloud.